Help advance science and conservation of butterflies
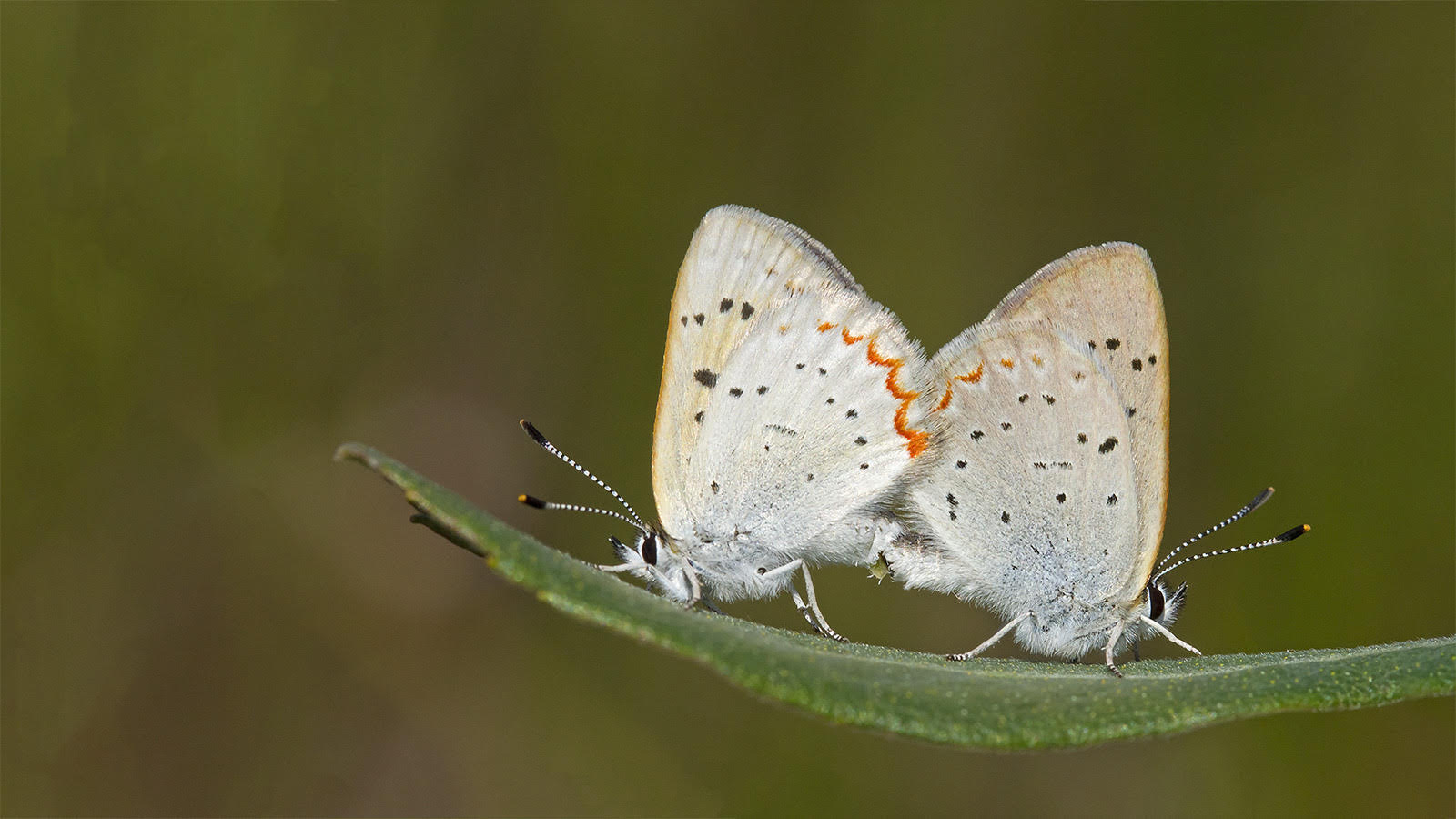
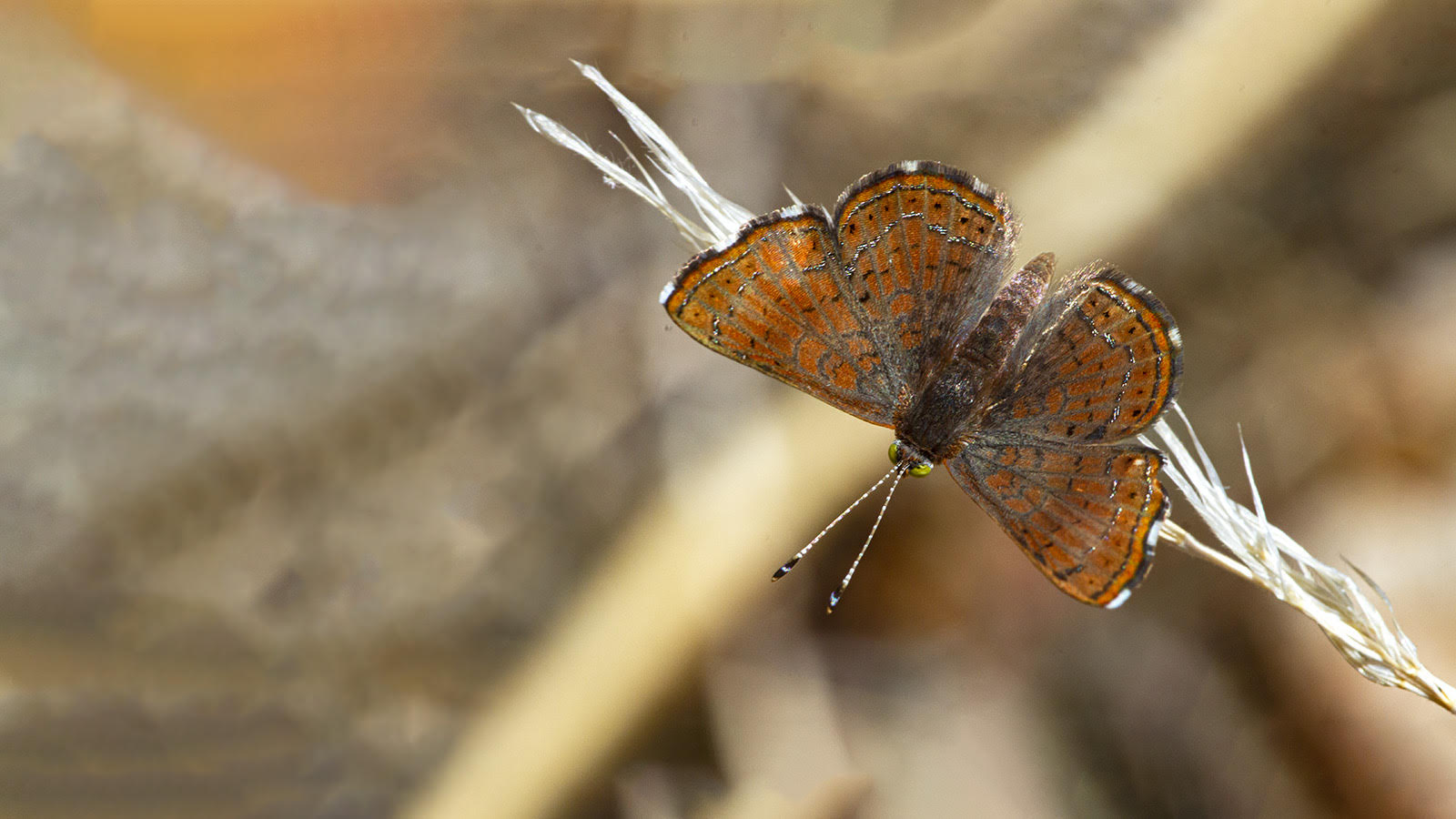
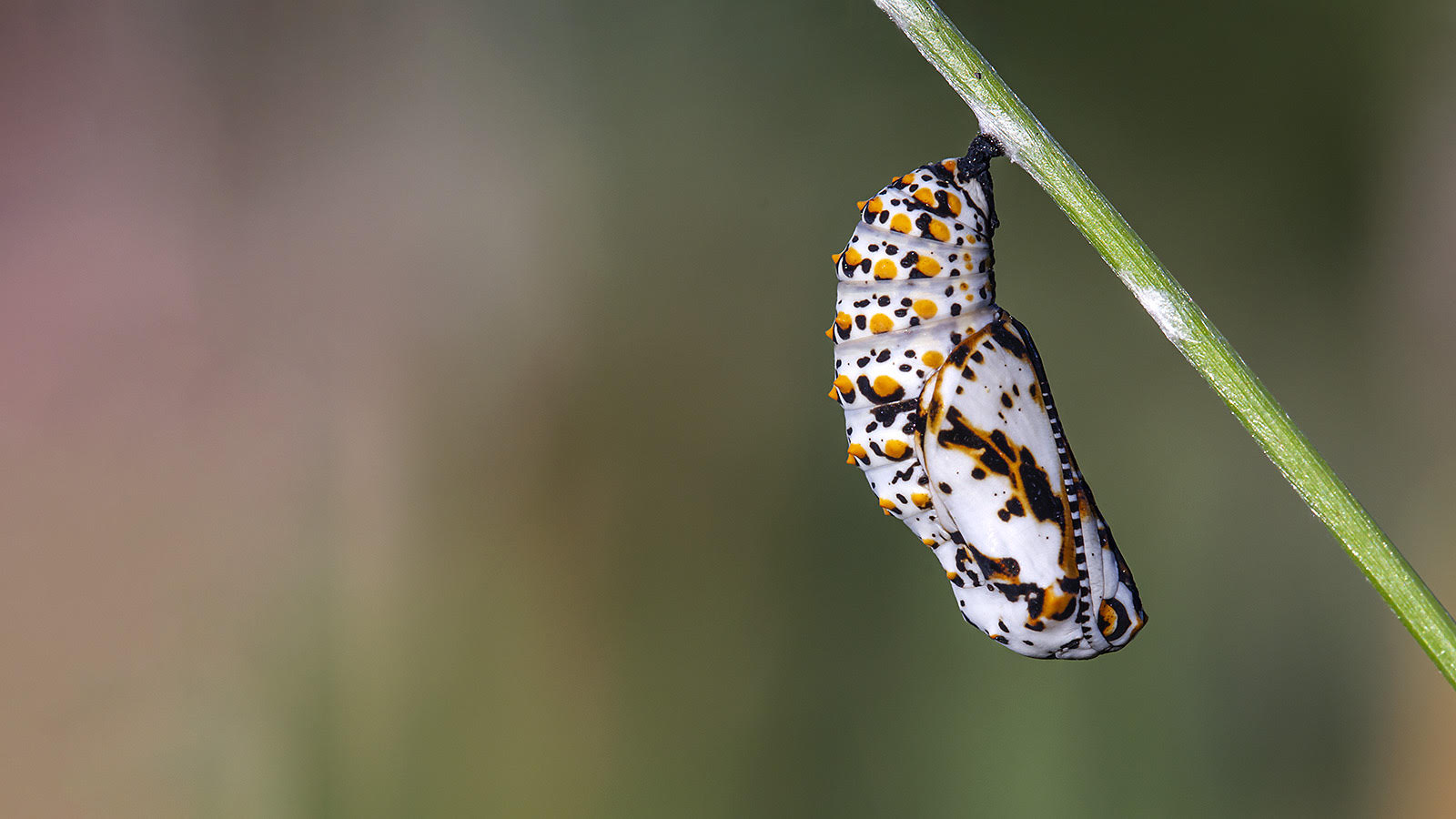
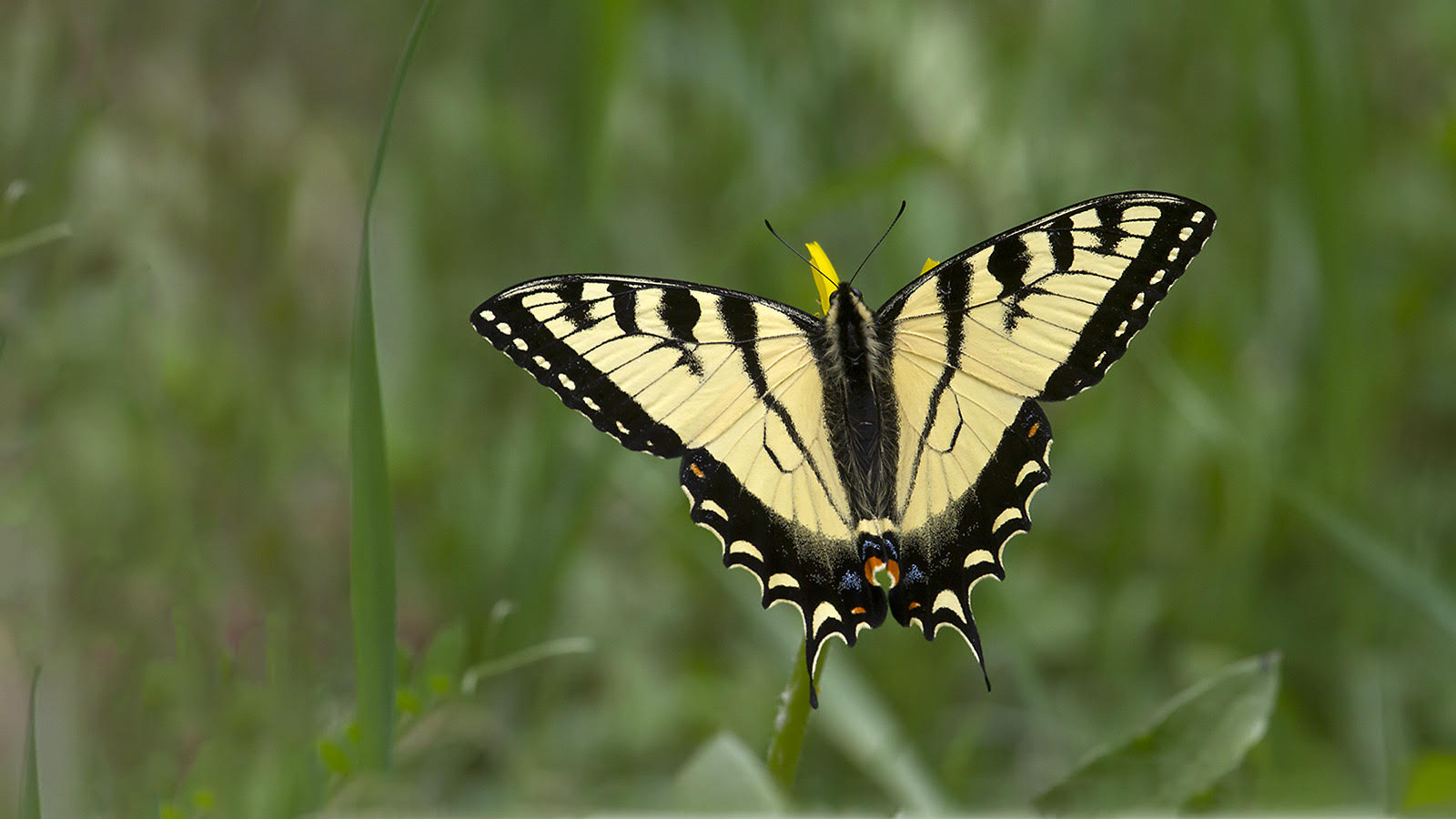
From the rarest butterflies to the most common, your sightings contribute to conservation decisions, scientific knowledge, education, and more. Help us understand when and where butterflies occur. All you have to do is watch and report your butterfly sightings.







